Sacrosanct Linux feature dies
I’ve been using Linux since 1996. Since then, the OS has undergone an amazing development, and all distributions provide an impressive high-resolution graphical interface. However, one feature has remained sacrosanct: the 6 virtual terminals. In the old days, when you had to provide timings for the video card and manually edit the xfree86 config file, it was easy to mess up the graphics display. But then, CTRL-Alt-F1 to the rescue! It was ALWAYS possible to get a terminal and consequently access to the operating system. And, in addition, most of the GUI versions of system setup programs had a TUI analogue, that could be run from the 80 x 24 terminal.
Until now. Ubuntu 12.04 BREAKS the virtual terminals on many older video cards, because it insists in using frame buffer mode, presumbly to provide fancy, meaningless, silly graphics for the boot screen. This is what my virtual terminals looks like now:
This is a scandal, no more, no less! Breaking the virtual terminals, that ALWAYS have been available, no matter what video card you had in your computer, breaks the promise that you always can obtain a console to control Linux. Simply a very, very bad design decision.
Python for crystallographers 5
Table of Contents
5 Python for crystallographers 5
This is the fifth and last in a series of tutorials on Python I gave at the Department of Molecular Biology, Aarhus University. It is aimed at crystallographers and structural biologists. This text is a commented transcript of the session log file produced by IPython The transcript does not include the output from Python. You will have to try it yourself.
5.1 Variable function arguments
Variable arguments can be used in a function when the number of arguments that the function needs to process cannot be predicted. The archetypical example is the C printf function, which accepts a format string as the first parameter, and any number of variables following that. In Python, variable function arguments are specified in the function definition by prefixing the name of the formal parameter with an asterisk:
def func(*args): print type (args) for i in args: print i,
From the point of view of the function, the passed parameters are contained in a tuple:
In [6]: func(1,2,'a') <type 'tuple'> 1 2 a In [7]: func(1,2,3,4,5,6) <type 'tuple'> 1 2 3 4 5 6
The function must of course know what to do with the varying number of arguments, and that is the job of the programmer.
5.2 Keyworded function arguments
This section introduces keyworded function arguments.
Keyworded arguments are somewhat similar to variable argument lists from the previous section, however, the function identifies the arguments by the parameter name, and arguments kan be skipped or they can be placed out of order. This is often a convenience to the programmer, and keyworded arguments are typically used to set various options in the function. Keyworded function arguments are specified in the function definition by prefixing the name of the formal parameter with a double asterisk:
def func1 (**kwargs): print type(kwargs) for k in kwargs: print k, kwargs[k],
From the point of view of the function, the passed parameters are contained in a dictionary. Let’s try to call this function using various keyworded arguments (recall that IPython’s %cpaste magic can be used to insert code snippets from the clipboard):
func1(x=1) func1(residues=100) func1(residues=100, atoms=1000) func1(atoms=1000,residues=100)
In [11]: func1(x=1) <type 'dict'> x 1 In [12]: func1(residues=100) <type 'dict'> residues 100 In [13]: func1(residues=100, atoms=1000) <type 'dict'> residues 100 atoms 1000 In [14]: func1(atoms=1000,residues=100) <type 'dict'> residues 100 atoms 1000
Quite typically, keyworded arguments are used to set optional parameters, but the function might still want one or more fixed and required parameters. A template example of such a use is given below:
def func2 (a, b, **kwargs): pass
and an example of a function call might look like this:
func2(filenam, 0.2, atoms=1000,residues=100)
5.3 Exceptions
Python is extremely smart about errors that arise during excecution of programs. When programs written in compiled languages such as C encounter an error, for example if a number is divided by zero, the operating system simply terminates the process. The programmer then needs to track down where the error happened, fix it, and recompile the program.
When running Python interactively, for example via IPython, the process survives, but it throws an exception, meaning that it stops what is was doing when the error happened. But since the process survives, you can examine the content of the variables, for example by printing them, and this can give vital clues about what went wrong. Of course, you can also use the Python debugger which streamlines this process, enables you to set breakpoints in the program and so on. Curiously to crystallographers, the Python debugger is called PDB. However, this subject is outside the scope of this presentation. IPython has a smart interface to the Python debugger, which you are encouraged to check out.
Here, we attempt to divide by zero:
In [15]: 1/0 --------------------------------------------------------------------------- ZeroDivisionError Traceback (most recent call last) /Users/mok/dropbox/python-course-2011/<ipython console> in <module>() ZeroDivisionError: integer division or modulo by zero
Python does not like that, and raises an exception named ZeroDivisionError. The programmer can also raise exceptions, both predefined, or home-made exceptions. Here, we try to raise at ZeroDivisionError:
In [16]: raise ZeroDivisionError --------------------------------------------------------------------------- ZeroDivisionError Traceback (most recent call last) /Users/mok/dropbox/python-course-2011/<ipython console> in <module>() ZeroDivisionError:
It is seen that an exception is thrown, but Python really doesn’t know the reason (there is no associated message.)
The very smart thing about exceptions is that you can catch them, and let the program deal with the unexpected situation. This is done in a
try/except block, for example:
a=10 b=2 try: f = a/b except ZeroDivisionError: print "oops!"
After the try/except block, everything is normal, and the value of f is 5. Let us try again, this time setting b to zero:
b = 0 try: f = a/b except: print "oops!"
The output written this time is “oops!”, and the value of f is unchanged (i.e. 5 if you follow the input above.)
5.3.1 The useful SystemExit exception
A quite common construction in a Python main program is to exit in the very beginning, if the user fails to provide the proper command line arguments, or, as in the following example, the specified file does not exist. When the SystemExit exception is raised, the Python process terminates:
fnam = sys.argv[1]
if not os.path.exists(fnam):
print "file not found, duh!"
raise SystemExit
5.4 Superposition
In this section, we will take a look at the Superimposer() class that comes with Biopython.
The following code is the main program from the Biopython source file Bio/PDB/Superimposer.py, that we copy-paste into its own file called superimposer.py. (The Python standard programming guidelines encourages use of lower-case letters for file names.)
1: import sys 2: from Bio.PDB import PDBParser, Selection, Superimposer 3: import numpy 4: 5: if __name__=="__main__": 6: 7: p=PDBParser() 8: s1=p.get_structure("FIXED", sys.argv[1]) 9: fixed=Selection.unfold_entities(s1, "A") 10: 11: s2=p.get_structure("MOVING", sys.argv[1]) 12: moving=Selection.unfold_entities(s2, "A") 13: 14: rot=numpy.identity(3).astype('f') 15: tran=numpy.array((1.0, 2.0, 3.0), 'f') 16: 17: for atom in moving: 18: atom.transform(rot, tran) 19: 20: sup=Superimposer() 21: 22: sup.set_atoms(fixed, moving) 23: 24: print sup.rotran 25: print sup.rms 26: 27: sup.apply(moving)
The above example is only mildly interesting, because it simply superimposes a molecule on a copy of itself, which has been translated by a small amount. The result of the calculation should therefore be the identity matrix, and a small translation in the opposite direction. We will none-the-less go through the program and examine what it’s doing.
The first few lines ought to be familiar by now. The PDB file from the command line is opened twice, and its atoms are placed in two Structure() instances s1 and s2.
In lines 9 and 12 we are using the convenient Selection() class, that can be used to access the various levels of the SMCRA hierachy. We use the method called unfold_entities(), which can unfold a list of entities to a list of entities of another level. For example:
- list of atoms -> list of residues
- list of residues -> list of atoms
- list of residues -> list of chains
In this case, we unfold a Structure() instance to a list of atoms, specified by the “A” in parameter 2. Those lists of atoms are stored in variables moving and fixed.
In line 14 we define the identity matrix, and in line 15 we define a translation vector (1.0, 2.0, 3.0). That rotation/translation operator is applied to the moving list of atoms in line 17, and after this operation all atoms in that list have been shifted by the translation vector given above.
Now we instance the Superimposer() class in the variable sup, and feed two lists of atoms to it (line 22). The rotation/translation operator and root mean squared deviation is printed, and finally, in line 27, the translation in sup is applied to the moving list, which should result in the atoms being shifted back to their original positions.
It is left as an exercise for the reader to generalize this program so it can take two different PDB files and/or two different chains.
5.5 Neighbor search
Now lets take a look at the NeighborSearch class from Biopython. This is a useful class that can search entities (that can be Atom, Residue, etc. objects) within a certain distance.
To remind yourself how smart IPython is (assuming you are running it) let us first check the documentation for NeighborSearch.
from Bio.PDB import NeigborSearch NeigborSearch?
Recall that IPython has extensive support of Tab-completion. When typing the above, it is actually sufficient to type:
Nei<TAB>?
and IPython will automatically expand “Nei” to “NeighborSearch”.
Now, load a Structure object from a PDB file, and as in the previous example, unfold that structure into a list of atoms.
from Bio.PDB import Selection s = p.get_structure("test", "1zen.pdb") atoms = Selection.unfold_entities(s, "A")
Now, choose a random atom in that structure. Below, we choose atom number 30, which happens to be the OH atom of residue 3, which is a Tyr. The get_coord() method returns a Numpy array containing that atom’s coordinates.
1: center = atoms[30].get_coord() 2: ns = NeighborSearch(atoms) 3: print ns.search (center, 5, "A") 4: print ns.search (center, 5, "R")
In line 2, we prime an instance ns of the NeighborSearch class with the list of atoms extracted from PDB structure 1zen. Then, in line 3, we print out the list of atoms within 5.0 Å of the center (atom OH of Tyr 3). In the following line, we print the list of residues found by the search within that radius:
Out[39]: [<Residue TYR het= resseq=3 icode= >, <Residue LYS het= resseq=105 icode= >, <Residue GLU het= resseq=53 icode= >, <Residue LYS het= resseq=52 icode= >]
NeighborSearch is a very useful class solving an often encountered problem, that you can use in your programs.
5.6 Fragment search
For the next example, we will take a look at Biopython’s Fragmentmapper class, and we shall use it to write a small program that performs a calculation more or less like the Lego_ca command in O: step through the polypeptide chain one residue at the time, and find the best fitting fragment from a library of fragments.
The FragmentMapper class requires that you download and install the fragment database first. It consists of a number of text files, that should be placed in a directory somewhere. You can download the fragment library from http://csb.stanford.edu/rachel/fragments/. In the following, it is assumed that these files are placed in a directory called fragment_data in the current working directory, but it could be anywhere on your file system. Here is the program fragmentmapper.py:
1: from Bio.PDB import PDBParser, FragmentMapper, Selection 2: import sys 3: 4: if __name__=="__main__": 5: 6: p=PDBParser() 7: s=p.get_structure("X", sys.argv[1]) 8: 9: m=s[0] 10: fm=FragmentMapper(m, 10, 5, "fragment_data") 11: 12: for r in Selection.unfold_entities(m, "R"): 13: print r, 14: if fm.has_key(r): 15: print fm[r] 16: else: 17: print
Most of the elements of this program should be familiar. The program accepts the file name of a PDB entry on the command line. In line 10, the FragmentMapper class is instanced. It receives a Model object, the size of the library (10 in this case), and the length of the fragments (we use 5 here, same as the Lego_* system from O). The last parameter is the directory where the fragment library files are stored.
In the loop starting at line 12 the model m is unfolded residue by residue, and each is looked up in the fragment mapper object, which functions like a Python dictionary. If there is a match, the fragment object is printed. The output of the program looks something like this:
<Residue SER het= resseq=1 icode= > <Residue LYS het= resseq=2 icode= > <Residue ILE het= resseq=3 icode= > <Fragment length=5 id=9> <Residue PHE het= resseq=4 icode= > <Fragment length=5 id=9> <Residue ASP het= resseq=5 icode= > <Fragment length=5 id=7> <Residue PHE het= resseq=6 icode= > <Fragment length=5 id=3> ...
The first two residues don’t have any matches, the first five-residue fragment match is at residue 3, because the position of the match is the central residue of the five. In other words, fragment id 9 is the best match of residues 1-5, and also the best match of residues 2-6, whereas the best match of residues 3-7 is fragment id 7.
5.7 Shell utilities
Finally, we will introduce a very useful module that is distributed with Python. It is the shutil module, containing various utility functions e.g. for copying files and directory trees.
import shutil dir(shutil)
Some of the functions available in shutil are:
- copy(src, dst)
- copy data and mode bits fra src to dst. Same as
“cp src dst” from the command line. - copy2(src, dst)
- copy data and all stat info (same as “cp -p src dst”).
- move(src, dst)
- Recursively move a file or directory to another location.
The shutil module is thus convenient for performing shell-like operations from within Python.
Date: 2012-02-01 12:08:44 CET
HTML generated by org-mode 6.21b in emacs 23
Python for crystallographers 4
Table of Contents
4 Python for crystallographers 4
This is the fourth in a series of tutorials on Python I gave at the Department of Molecular Biology, Aarhus University. It is aimed at crystallographers and structural biologists. This text is a commented transcript of the session log file produced by IPython The transcript does not include the output from Python. You will have to try it yourself.
4.1 Python modules
We have seen that simple Python modules are often just files containing Python code, typically functions and classes. In presentation 2, we saw the following construct:
from utils import parse_list
that simply imports the function parse_list() from the file utils.py. This allows the programmer to organize the code in elements that are more easily maintained. However, many large projects are exported as packages. A package is normally a collection of modules, but can also be a single module.
To define a module called course, we can create a directory called course, and in that directory, we place a file called __init__.py. The presence of this file in a directory tells Python that it is a module.
mkdir course/ touch course/__init__.py
Now, inside Python, we can:
import course
but of course, the module is empty. There are several ways to put code into a module. Most often, __init__.py is empty, but it can contain Python code. Let us enter the following into the file course/__init__.py:
def hello(): print "Python is cool"
Now, in IPython, this function can be called:
import course
course.hello()
You can see that the function hello() is now in the modules’ namespace. The directory course can also contain other Python files, which then become part of the module. Let us copy our file from earlier utils.py to the new course module:
cp utils.py course/
Now, utils.py is a part of the course module, and needs to be addressed as a part of that. In Python, we can import the function
parse_list() like this:
from course.utils import parse_list
This imports the function parse_list() directly into the current namespace. However, we can also write:
import course
but then we need to use the fully qualified namespace to access the function which should be called course.utils.parse_list(). It is a matter of taste, and a matter of code clarity, how to use the import statement in your program. As a rule-of-thumb, if you need to use the function many places in your code, it is convinient to import its name into the current namespace. If you only need to call a function (or class) once, it is more clear to leave it fully qualified, because the code then will show its origin.
4.2 The Bio.PDB module
We will now look at the PDB module from Biopython. This module is quite sophisticated, and is supposedely be very well tested. Basically, the PDB module regards a macromolecular structure from a PDB entry as a hierachy called “SMCRA”, which stands for
- Structure
- Model
- Chain
- Residue
- Atom
In a typical X-ray structure, the model level is not used, but in a typical NMR structure entry, it is typical to have 20 models or more in one file. Each Model contains one or more chains, each Chain contains one or more residues, and each Residue contains one or more atoms.
The PDB module is designed so it is possible to access this information in several ways. How, exactly, this is done in your program depends on the requirements of the specific application.
4.2.1 Parsing a PDB file
To parse a PDB file we need to instantiate a parser. The parser object has a few methods to access the content of the PDB file, but one method is always used, namely get_structure() which – as the name implies – returns a Structure object.
1: from Bio import PDB 2: 3: p = PDB.PDBParser() 4: s = p.get_structure("3ns0", "3ns0.pdb")
In line 3 the parser object is instantiated in the object
p. Next, in line 4 we retrieve the Structure object. The method get_structure() needs an identifier string (can be any meaningful string, here we use the PDB ident) and the file name of a PDB entry.
In fact, the second argument to get_structure() can also be a file handle, or generally, an object that has a readlines() method. This is useful if for example you would like to read a gzipped PDB entry:
import gzip fd = gzip.open("1psr.pdb.gz") s1 = p.get_structure("1psr", fd)
The gzip module transparently opens both gzipped and text files.
4.2.2 The Structure object
The Structure object is the top-level object, and contains methods to access the child objects. Many of these methods are in fact common to the lower-level objects also, because they are inherited from a common base class called Entity(). The methods include:
get_level()- return level in hierarchy (“A”, “R”, “C”, “M” or “S”)
get_list()- return list of children
get_id()- return the ID of the object
The objects in the SMCRA hierachy also inherit attributes from the base class, including child_dict, which is a dictionary containing the child objects with the child IDs as keys.
However, the Structure object also contains convienient methods that more directly access the content of the structure:
get_chains()- return a chain iterator
get_residues()- return a residues iterator
get_atoms()- return an atom iterator
Recall that iterators are objects that cannot be accessed directly,
but must be used in a loop contruct.
Finally, the Entity() base class defines the special method __getitem()= which enables the use of the “[]” operator on the object. In our example, s[0] thus contains the child (a Model object) with ID 0. Another usefule attribute is parent, which contains a reference to the parent object.
4.2.3 The Model object
The Model object is used in PDB entries containing NMR models. An NMR entry typically contains 20 models or more. PDB entries of structures determined by X-Ray crystallography normally only has one model.
4.2.4 The Chain object
Each Model object contains a list of one or more Chain objects. One Chain object is generated from each unique chain identifier in the PDB file (column 5), and the Chain object controls access to the residues that are a part of that chain.
4.2.5 The Residue object
The pattern should be obvious by now. The Residue object defines one residue in the structure.
To retrieve residues in a structure, we first store the model in object m, then loop over chains c, and for each chain, we can loop over residues r:
m = s[0] for c in m: for r in c: print r
However, there really is no need to loop over models, then chains, then residues unless you actually need to examine the structure in this way. We can use a convienience method in the Structure object to retrieve all residues from a structure directly:
1: R = s.get_residues() 2: for r in R: 3: print r.id, r.parent.id
Line 1 gives us an iterator that can be used to visit all residues belonging to structure s. In line 3 we print the ids of all residues. This is the number given in column 6 of a PDB file. Many molecular graphics programs prefix the residue number with the chain ID (for example “A129”) but here, we retrieve the ID of the parent object (the parent of Residue objects is the Chain).
4.2.6 The Atom object
The Atom() class is the most fundamental class in the SMCRA hierachy, and also the class with the most methods. In particular, we have the get_coord() method, which returns the coordinates of the atom in a Numpy array. We can retrieve atoms from a structure using the get_atoms() method:
4: A = s.get_atoms() # get an atom iterator 5: L = [] 6: for a in A: 7: L.append(a.get_coord()) 8: print L
In line 7, we stash away the coordinates of a structure in a list L. The Atom object also exposes methods to retrieve the B-factor, the occupancy, serial number, etc.
4.3 List comprehension
In this and earlier presentations, you have seen that we routinely extract objects and organize them into a list. For example in the example above we first created an empty list L, and in the following loop, we extract the objects we are interested in and append them to the list. This construction is used to often in Python, that a special idiom in the language has been created to deal with it. It is called list comprehension. Let us revisit the case of retrieving atoms from a structure:
A = s.get_atoms() # get an atom iterator L = [a for a in A]
This construction might look a bit weird when you first see it, but it quickly becomes a part of your Python vocabulary. Let us break the syntax down a bit. The initial a is the object that gets appended to the list, or in other words, the list will consist of a‘s. The following “for” statement determines what a‘s will be selected, namely those generated by the iterator A.
We can condense it even further:
L = [a for a in s.get_atoms()]
This gives us a list of atoms in L. However, in the example in the previous section (7) we retrieved a list of coordinates not atoms. Using a list comprehension, we can write:
L = [a.get_coord() for a in s.get_atoms()]
A list comprehension can also introduce an if-filter, for example:
T = [r for r in s.get_residues() if r.resname=='TYR']
creates a list T containing tyrosine residues.
4.4 Distance matrix plot
We will now use Biopython and Numpy to write a program to make a C-alpha distance matrix plot. The core logic of this program looks like this:
1: s = parser.get_structure(id, fnam) 2: 3: chains = [c for c in s.get_chains()] 4: 5: x = [] 6: for r in chains[0]: 7: if 'CA' in r.child_dict: 8: ca = r.child_dict['CA'] 9: x.append(ca.get_coord())
First, we generate a list of chains (line 3), and then, in the loop, we step through the residues of the first chain only. It is left as an exercise to make the program more general so that it can deal with all chains, or deal with a specific chain specified by the user on the command line. In line 7, we check to see if the key ‘CA’ is in the residues’ child dictionary. For a residue, this dictionary contains the atoms of the residue, indexed by their names. If the atom has the name ‘CA’, the atom object is retrieved, and its coordinate appended to the list x.
The next step is to convert the list of coordinates (each of which is a Numpy array) to a large Numpy array of dimensions (N,3) where N is the number of atoms:
coords = np.array(x)
The final step is to compute the distance matrix. Fortunately, this is possible by using a module from scipy. The function cdist is able to compute a distance matrix between two different sets of coordinates, in our case, we need to compute a distance matrix between the coordinates to themselves. This is why the first two arguments passed to cdist are both coords. The last argument specifies that we want to compute the eucledian distance (normal geometry).
import scipy.spatial data = scipy.spatial.distance.cdist(coords, coords, 'euclidean')
Now, with the distance matrix stored in the Numpy array data, all we need to do is plot it:
plot (data)
The code in the plot() function was copied from the Gallery on the matplotlib web site, after first finding a plot that looks like the one we want.
The entire source of the program distancematrix.py, including the function plot() is shown in the Appendix.
Appendix
The source code of the program distancematrix.py. In the plot() function, the commented-out code is needed to create a plot to a PNG file instead of to the computer screen. To generate a plot, the lines:
# import matplotlib # matplotlib.use( 'Agg' )
should be activated, and the function pylab.savefig() should be used instead of pylab.show().
import sys import os from Bio.PDB.PDBParser import PDBParser import numpy as np import warnings def plot(data): # import matplotlib # matplotlib.use( 'Agg' ) import pylab fig = pylab.figure() ax = fig.add_subplot(111) cax = ax.imshow(data, interpolation='nearest') ax.set_title('Ca-Ca distance plot') # Add colorbar, make sure to specify tick locations to match desired ticklabels min = np.min(data) max = np.max(data) cbar = fig.colorbar(cax, ticks=[min, max]) cbar.set_ticks([min,max]) pylab.show() # pylab.savefig( 'distmat.png', format='png' ) if __name__ == '__main__': import re fnam = sys.argv[1] if not os.path.exists(fnam): print "file not found, duh!" raise SystemExit id = fnam.split('.')[0] warnings.simplefilter('ignore') parser = PDBParser() s = parser.get_structure(id, fnam) chains = [c for c in s.get_chains()] x = [] for r in chains[0]: if 'CA' in r.child_dict: ca = r.child_dict['CA'] x.append(ca.get_coord()) coords = np.array(x) import scipy.spatial data = scipy.spatial.distance.cdist(coords, coords, 'euclidean') print data.shape plot(data)
Date: 2012-01-30 13:59:32 CET
HTML generated by org-mode 6.21b in emacs 23
Python for crystallographers 3
Table of Contents
3 Python for crystallographers 3
This is the third in a series of tutorials on Python I gave at the Department of Molecular Biology, Aarhus University. It is aimed at crystallographers and structural biologists. This text is a commented transcript of the session log file produced by IPython The transcript does not include the output from Python. You will have to try it yourself.
3.1 Reading a PDB file
The first project is to read a PDB format file. The following snippet of code opens a file and swallows it. The list L will contain a list of lines:
fnam = '1au1.pdb' f = open(fnam) L = f.readlines() print L[10]
But L includes a lot of header lines from the PDB file that we really don’t need. To help filter out the ATOM records, we make use of Pythons regular expression module, called re. Then we step through all lines in the list L, extract the lines that start with ‘ATOM’, and append those to another list: data.
import re data = [] for rec in L: if re.match('ATOM', rec): data.append(rec.split())
We check the length of the list data, and print out a few of the first atoms:
len(data) data[0] for i in 1,2,3,4: print data[i]
Remember, that Python arrays start at 0, so data[0] contains the first atom, but that atom has serial ID number 1.
3.2 Class based approach
Now we will create a more advanced and flexible data structure for atoms. We create a file pdb.py with the following content:
1: class Atom: 2: def __init__(self): 3: self.name = '' 4: self.id = 0 5: self.element_number = 6 6: self.x = 0.0 7: self.y = 0.0 8: self.z = 0.0 9: self.b = 1.0 10: self.occ = 1.0
In line 2 we define the class constructor. In this design, the constructor does not need any parameters. So now we can define an “empty” atom. In IPython, we use the %run magic command to “source” the file pdb.py:
%run pdb atom = Atom() print a.x, a.y, a.z, a.occ, a.b
This will print out the default values defined in the constructor.
Now, let us add a method to the Atom class that will parse an ATOM record from a PDB file, and fill the relevant attributes with the relevant data. Here is the method, it should be addede to the class Atom in the file pdb.py:
def parse_from_pdb (self,s): L = s.split() self.id = int(L[1]) self.name = L[2] self.x = float(L[6]) self.y = float(L[7]) self.z = float(L[8]) self.occ = float(L[9]) self.b = float(L[10])
Now, in IPython, we run again:
%run pdb
atom = Atom()
We still have a record from the PDB file from before, when we loaded the PDB file into a list of strings called L. Let us try to parse one of these lines:
atom.parse_from_pdb(L[2000])
print atom.x, atom.y, atom.z
This loaded the data from line 2000 in the file into the object atom.
3.3 version 2
Now we would like to make a more complete standalone program, so we add a main program to the file pdb.py:
1: class Atom: 2: def __init__(self): 3: self.name = '' 4: self.id = 0 5: self.element_number = 6 6: self.x = 0.0 7: self.y = 0.0 8: self.z = 0.0 9: self.b = 1.0 10: self.occ = 1.0 11: 12: def parse_from_pdb (self,s): 13: L = s.split() 14: self.id = int(L[1]) 15: self.name = L[2] 16: self.x = float(L[6]) 17: self.y = float(L[7]) 18: self.z = float(L[8]) 19: self.occ = float(L[9]) 20: self.b = float(L[10]) 21: #. 22: 23: if __name__ == '__main__': 24: import re 25: 26: fnam = '1au1.pdb' 27: f = open(fnam) 28: L = f.readlines() 29: 30: atoms = [] 31: for line in L: 32: if re.match('ATOM|HETATM', line): 33: a = Atom() 34: a.parse_from_pdb(line) 35: atoms.append(a)
We open the file as before, and swallow the whole thing using readlines(). This gives a list of lines. In line 30, we create an empty list that will be used to store the atom objects. In line 31 we step through all the strings in list L. Something new happens in line 32. Here we use match() of the regular expression module re to “grep” out lines that start with either ‘ATOM’ or ‘HETATM’. Only if one of these two keywords are found, we will execute the code inside that if statement. If we have an ‘ATOM’ or a ‘HETATM’ record, we instance an empty Atom object, and use the parse_from_pdb() method to populate the fields of the object. Finally, we append the newly created object to the list atoms. Now, run this from IPython, and look at some of the elements in list atoms:
%run pdb print atoms[1] print atoms[10]
This printout is pretty uninformative. We should add a __repr()__ method to the Atom class. It looks like this:
def __repr__(self): s = "Atom: {0}, x,y,z={1},{2},{3}, occ={4}, B={5}" s = s.format(self.name,self.x,self.y,self.z,self.occ,self.b) return s
Here, the format() method of the str class is used to format the string. In this case, the tokens marked with curly brackets will be formatted with the corresponding argument. There is a whole mini-language that allows very sophisticated string formatting, it can be studied here: http://docs.python.org/library/stdtypes.html#string-formatting
%run pdb
print atoms[1]
which prints output that looks like this:
Atom: CA, x,y,z=24.887,27.143,6.222, occ=1.0, B=41.36
Now, we can enhance our main program to write out more information:
if __name__ == '__main__': import re fnam = '1au1.pdb' f = open(fnam) L = f.readlines() atoms = [] for line in L: if re.match('ATOM|HETATM', line): a = Atom() a.parse_from_pdb(line) atoms.append(a) for a in atoms: print a
However, when we run this, we get a run-time error!
In [7]: %run pdb 1au1.pdb
---------------------------------------------------------------------------
ValueError Traceback (most recent call
last)
/u/mok/Dropbox/python-course-2011/pdb.py in <module>()
59 if re.match('ATOM|HETATM', line):
60 a = Atom()
---> 61 a.parse_from_pdb(line)
62 atoms.append(a)
63
/u/mok/Dropbox/python-course-2011/pdb.py in parse_from_pdb(self, s)
18 self.y = float(L[7])
19 self.z = float(L[8])
---> 20 self.occ = float(L[9])
21 self.b = float(L[10])
22 #.
ValueError: invalid literal for float(): 1.00100.00
WARNING: Failure executing file: <pdb.py>
The second-last line gives us a hint to what is going on. Python can not convert the string ‘1.00100.00’ to a floating point number. This is a well known limitation of the PDB format. When a B factor becomes 100.0 or larger, there is no longer a space between the occupancy field and the B-factor field. Therefore, the logic we used in the parse_from_pdb() method is too simplistic. We can’t simply split the line into space-separated fields using the split() method. We need to be more careful. So, parse_from_pdb() needs to be changed to this:
def parse_from_pdb (self,s): L = s.split() self.id = int(L[1]) self.name = L[2] self.x = float(L[6]) self.y = float(L[7]) self.z = float(L[8]) self.occ = float(s[55:60]) self.b = float(s[60:66])
Now, instead of using the split fields to extract occ and b, we extract them directly from positions 55 to 59 and 60 to 65 in the input string. (This will work for this file, but the logic is likely to fail if tested on every PDB file we can find, because there are other issues with PDB files that may cause the number of fields to be different.)
3.4 A more complete program
Now, let us make the program more versatile, so that we can specify the PDB file from the command line. We change the main program to look like this:
1: if __name__ == '__main__': 2: import re 3: import sys 4: import os 5: 6: fnam = sys.argv[1] 7: if not os.path.exists(fnam): 8: print "File", fnam, "not found, duh!" 9: raise SystemExit (ref:exit) 10: 11: f = open(fnam) 12: L = f.readlines() 13: 14: atoms = [] 15: for line in L: 16: if re.match('ATOM|HETATM', line): 17: a = Atom() 18: a.parse_from_pdb(line) 19: atoms.append(a) 20: 21: for a in atoms: 22: print a
In line 6 we use the sys module to access the (first) command line argument. This is assumed to be a file name, but in line 7 we double check to make sure the file exists, by using exists() from the (extremely useful) os.path module. If the file is not found, we print out an error message, and stop the program by raising an exception in line nil. Now the program is more general, and can be used to read in any PDB file.
3.5 Distance between atoms
We want to be able to compute the distance between two atoms. We add a function to pdb.py to do this. The function assumes to be passed two objects that have attributes x, y and z, it then does the math computes the distance between these points.
def distance(atom1, atom2): import math dist = math.sqrt((atom1.x-atom2.x)**2 +(atom1.y-atom2.y)**2 +(atom1.z-atom2.z)**2) return dist
We need to import the math module so we can look up the square root. We could also have written the function as a method of the Atom class, in which case the call would be:
d = atom1.distance(atom2) print d
This is a matter of design. We choose to write a function, that is called like this:
d = distance(atom1, atom2) print d
We can then compute the distance of all atoms to (a random) atom number 300 The full source code of the program is listed in the appendix. We run the program from within IPython like this:
%run pdb 1au1.pdb
3.6 Difference between = and ==
The answer is simple: = is used for assignement, and == is used for comparisons. For example:
In [9]: a=1 In [10]: a Out[10]: 1 In [11]: a == 2 Out[11]: False In [12]: a == 1 Out[12]: True
3.7 Plotting with matplotlib
Finally, let us take a look at the incredibly useful plotting library, matplotlib, that is a part of SciPy. First, we create a list of B values from our list of atom objects:
bvalues =[] for a in atoms: bvalues.append(a.b)
Now we can make plot of B-values vs. atom number:
import pylab
pylab.plot(bvalues)
Matplotlib can do a huge amount of different plots. A great way to get started is to go to the matplotlib gallery at http://matplotlib.sourceforge.net/gallery.html and choose a plot that looks like what you need. Then you can cut and paste the source code of the plot into IPython. Use the %cpaste magic to do this.
Appendix
Complete source code of the final version of pdb.py
class Atom: def __init__(self): self.name = '' self.id = 0 self.element_number = 6 self.x = 0.0 self.y = 0.0 self.z = 0.0 self.b = 1.0 self.occ = 1.0 def parse_from_pdb (self,s): L = s.split() self.id = int(L[1]) self.name = L[2] self.x = float(L[6]) self.y = float(L[7]) self.z = float(L[8]) self.occ = float(s[55:60]) self.b = float(s[60:66]) def __repr__(self): s = "Atom: {0}, x,y,z={1},{2},{3}, occ={4}, B={5}" s = s.format(self.name,self.x,self.y,self.z,self.occ,self.b) return s def distance(atom1, atom2): import math dist = math.sqrt((atom1.x-atom2.x)**2 +(atom1.y-atom2.y)**2 +(atom1.z-atom2.z)**2) return dist if __name__ == '__main__': import re import sys import os fnam = sys.argv[1] if not os.path.exists(fnam): print "file not found, duh!" raise SystemExit f = open(fnam) L = f.readlines() atoms = [] for line in L: if re.match('ATOM|HETATM', line): a = Atom() a.parse_from_pdb(line) atoms.append(a) XXX = atoms[299] for a in atoms: print distance(a,XXX)
Date: 2012-01-29 16:22:46 CET
HTML generated by org-mode 6.21b in emacs 23
Oneiric on the Dell Mini 10: Nope
A couple of months ago, I downloaded the Ubuntu 11.10 installer and put it on an USB stick. Judging from the discussions on the net, I might be one of the only people on the planet who actually *like* Unity, and I was really looking forward to getting it on my netbook. In my opinion, Oneiric Ocelot is a terrific release. I have it on my workstation and I like it a lot.
Unfortunately, the install on my miserable Dell Mini 10 failed. It dies and freezes. Frustration. I wanted to blog about it, but didn’t get around it until now. So here is what happens. The following screenshots were taken with my smartphone, that is the only way I could make screendumps.
Shortly after the Ubuntu 11.10 screen, with the five dots, the following appears:
After some seconds the purple background color turns black:
… and then the booting process shuts down:
I have no idea what’s going on, but it seems to have something to do with the wireless driver. Next thing to try is to boot while the Dell Mini 10 is wired to the network.
Python for crystallographers 2
Table of Contents
2 Python for Crystallographers 2
This is the second in a series of tutorials on Python I gave at the Department of Molecular Biology, Aarhus University in 2011. It is aimed at crystallographers and structural biologists. This text is a commented transcript of the session log file produced by IPython The transcript does not include the output from Python. You will have to try it yourself.
2.1 Classes
Classes are a fundamental feature in all Object Oriented programming languages. C++, Java, Objective-C, Smalltalk, etc. all implement classes. Programmers use classes in different ways. Some classes represent concrete objects, for example atoms, residues, or sequences, but in other cases, classes are used to encapsulate various algorithms, or they produce iterators or are factories of objects. As you use various Python modules, you will meet different class designs.
Let us define a very simple class:
class A: a = 1
dir(A)
A Python class is not usually used on its own. You can compare a class to a cookie-cutter, it defines the shape and size of the cookie, but to actually get something concrete out of it, you need to press it into the dough to make a cookie. With classes, you need to instance it to make it useful, and you can create as many instances as you want (unless the class has specific logic to prevent that.)
inst = A()
inst.a
Class A only contains one variable, a. Variables of this kind are called class variables, because they are shared by all instances of the class. This is useful for e.g. constants and other data items, but in general, instances need their own data too.
Here we define a class that has an instance variable b:
class A: a = 1 def __init__(self, name): self.name = name
The special method __init__()= is called whenever Python constructs an instance of a class. The variable self refers to the instance object itself, so self.name refers to a variable in the namespace of the instance. The __init__() method can also be called a constructor. When we define a constructor that takes an argument, the instance must be defined by passing one to the constructor. This fails:
inst = A()
but this works:
inst = A("abc") print inst.name
We can make another instance of class A:
inst2 = A("def") print inst2.a print inst2.name
You see that the name instance variable is different in the objects inst and inst2, but the class variable a is the same. If we change a:
A.a = 2 print inst2.a print inst.a
You will see that the class variable a has now been set to 2 in both instances.
2.1.1 Class inheritance
Now we look at class inheriance. Here is a class that describes a person:
class Person: def __init__(self, name, addr, email, phone): self.name = name self.address = addr self.email = email self.phone = phone def set_email(self, email): self.email = email print "email sat til", email
The instance variables name, address, email, and phone is defined in the constructor, so it is used like this:
pers1 = Person("morten", "århus", "mok@example.com", 86188180)
We can access the attributes via the instance of the person:
print pers1.name print pers1.email
In the class definition, we have defined a method to set the email address. A method of this type is called a “setter”:
pers1.set_email('nogetandet@gmail.com')
In this case, the setter does nothing else than change the value of the email variable, so we could have done just this:
pers1.email = 'nogetandet@gmail.com'
however, using getters and setters is normally considered good programming style.
If we want to have a class describing a student, we can make use of the functionality already existing in the Person class. This is called the DRY principle: Don’t Repeat Yourself. A student however, has special attributes that is not used for other persons:
class Student(Person): def set_studiekortnummer(self, s): self.studie_nr = s
We have added a method to set the studynumber, but the name, address etc. is inherited from the base class.
rune = Student("rune", "ukendt", "rune@gmail", 89421111)
We now have an instance of the Student class called rune. Let us set the student number:
print rune rune.set_studiekortnummer(123456) print rune.studie_nr print rune.name
2.2 Biopython
We will now look at the Biopython package, but first, let us write a small program to fetch a sequence from Uniprot that is purely native Python.
2.2.1 fetch sequence from Uniprot
Here is the content of the program called fetch-files-uniprot.py:
#! /usr/bin/env python from utils import parse_list import urllib import sys uniprot = "http://www.uniprot.org/uniprot/" remote = urllib.FancyURLopener() if __name__ == "__main__": fd = open (sys.argv[1]) for ent in parse_list(fd): print "getting ", ent f = remote.open(uniprot+ent+".txt") seq = f.read() f.close() g = open(ent+".txt", "w") g.write(seq) g.close() #. #.
Let us examine the code. Near the top, you see that we import the function parse_list from the module utils. This is our own Python module, stored in a file utils.py, and it looks like this:
def parse_list(f): "Parse our little sequence id list files" line = 'begin' L = [] for line in f: if '#' in line: line,comment = line.split('#',1) #. if len(line) > 0: L.append(line.strip()) #. return L #.
This function reads a file, skips comments marked with ‘#’, and returns a list of sequence IDs.
Next, we open a file with the file name sys.argv[1]. This is an attribute from the imported sys module, that contains the parameters given on the command line. So, it our program is invoked like this:
$ python fetch-files-uniprot.py ifns.txt
the value of sys.args[1] will be ‘ifns.txt’. Our program then proceeds by reading entries one by one from the file-like object remote, which is an instance of the object urllib.FancyURLopener() from the urllib module, opened with the URL of the relevant sequence entry. Every sequence entry is then stored in a local file.
2.2.2 Fetch sequence using Bio.Entrez
Now we will fetch some sequences using NCBI’s Entrez database. Biopython provides a convienient interface to Entrez’ API. Here is the program fetch-files-entrez.py:
#! /usr/bin/env python from Bio import Entrez from utils import parse_list import sys # See http://www.ncbi.nlm.nih.gov/entrez/query/static/efetchseq_help.html file_ext = {'gp':'gb', 'fasta':'fa', 'ft':'ft', 'gpc':'gpc', 'full':'full', } if __name__ == '__main__': fd = open (sys.argv[1]) Entrez.email = "mok+entrez@example.com" for ent in parse_list(fd): for rettype in file_ext: handle = Entrez.efetch(db="protein", id=ent, rettype=rettype, retmode="text") fnam = ent + '.' + file_ext[rettype] g = open(fnam, 'w') g.write(handle.read()) g.close() print "wrote", fnam #. #. #.
This program is a bit more complicated than the Uniprot fetching program, but follows the same general logic. First, we set a class variable Entrez.email, this is requested by the Entrez service and we do that as a matter of courtesy. As before, we pass through all sequence IDs in the input file, but this time, we add an additional loop inside, that loops through a list of different data return types given in the dictionary file_ext, which at the same time maps a file extension to each return type. After running this program, we are left with a series of files for each sequence ID, for example, for ID P01563 we have:
-rw-r--r-- 1 mok mok 345 May 30 10:11 P01563.fa -rw-r--r-- 1 mok mok 2676 May 30 10:11 P01563.ft -rw-r--r-- 1 mok mok 80468 May 30 10:11 P01563.full -rw-r--r-- 1 mok mok 17099 May 30 10:11 P01563.gb -rw-r--r-- 1 mok mok 56359 May 30 10:11 P01563.gpc -rw-r--r-- 1 mok mok 11662 May 30 10:03 P01563.txt
Now, lets go back to the IPython interface and play a bit more with Biopython. First, we will use the SeqIO module:
from Bio import SeqIO dir (SeqIO) # look at what SeqIO contains seq = SeqIO.read("P01563.gb", "genbank")
Now, we have a Biopython sequence object (a “SeqRecord”) named seq containing all the information that was parsed from the input Genbank format file:
dir(seq) print(seq)
We can look at the sequence description, and we see that the sequence is stored in a special object which also contains the alphabet.
seq.description seq.seq
The object seq can be indexed using the square bracket notation (like normal Python lists), and it can be iterated in a loop:
print seq[0] for a in seq: print a
The SeqRecord object attribute features contains a list of feature objects, all collected from the Genbank format sequence file:
print seq.features
You can investigate this yourself more, look at the various types of features and what data they contain.
2.3 The special method __repr__
We will briefly look at the special method called __repr()__. This is a method that can be defined in any class, and it defines how the class behaves when passed to the print function, and should simply return a string representation of the object. Let us define this method in our Person class from before:
class Person: def __init__(self, a, addr, email, tlf): self.name = a self.adresse = addr self.email = email self.telefon = tlf def set_email(self, email): self.email = email print "email set to", email def __repr__(self): return "Name: " + self.name
Now, if we print the person pers1 from before, Python will print:
Name: morten
instead of a strange technical object description. As the base class is now enhanced, the new capability will automatically be available to the Student class too:
rune = Student("rune", "ukendt", "rune@gmail", 89421111) print rune
2.4 Function returning tuples
A function can return a tuple:
def f(): return "a", "b" print f() i,j = f() print i print j
The above example shows how each element in the tuple returned by function f() can be assigned to different variables i and j.
2.5 Listdir
Python is very suited for all kinds of systems work. The os module contains lots of useful functions for interacting with the operating system. For example, the function listdir() will return a list of files in the directory passed as an argument:
import os this_dir = os.listdir('.') tmp_dir = os.listdir('/tmp') for f in os.listdir('/tmp'): print f
Date: 2012-01-26 18:53:42 CET
HTML generated by org-mode 6.21b in emacs 23
Python for crystallographers 1
Table of Contents
1 Python for Crystallographers 1
This is the first in a series of tutorials on Python I gave at the Department of Molecular Biology, Aarhus University in 2011. It is aimed at crystallographers and structural biologists. This text is a commented transcript of the session log file produced by IPython The transcript does not include the output from Python. You will have to try it yourself.
1.1 IPython
Python has a quite decent interactive mode that offers readline editing of your input. However, IPython is an extension of Python that offers some very convenient features. IPython has a number of so-called “magic” commands that can be listed using
%lsmagic
Magic commands start with the percent sign, but if the automagic mode is activated (IPython uses that mode by default) you can omit it. I will activate logging of my session using the magic:
logstart
There are several very good screencasts for IPython at http://showmedo.com, search for tutorials by Jeff Rush and unpingco. You will see that you can do some very cool stuff with IPython!
1.2 Simple Python variables
Python has simple variables, integers, floats, complex and strings. Here is an integer:
a = 1
and a float:
f = 1.
We can see what variables we have assigned with the magic commands %who and %whos. The latter offers more detail.
who
Strings are defined by strings surrounded by quotes or double quotes, so “python” and ‘python’ is the same thing. If you need to put quotes or double quotes inside a string variable, you can do that like this:
s = '"python"' s = "python's way"
Complex numbers are defined like this:
b = complex(1,5)
A complex number has real and imaginary parts. These are stored in the object attributes real and imag:
b.imag b.real
A complex number can be used in arithmetic expressions as expected:
a = 3.4
b*a
b*b
1.3 Compound objects
1.3.1 Lists
Pythons true strength comes from the powerful built-in compound objects. First, we have the list, which is defined using square brackets.
L = []
This defines an empty list. When running interactively, you can inspect the value of objects by typing their name,
L
but when running a Python program, you need to print it:
print(L)
Here is a list containing integers:
L = [2,4,6,8]
We can append numbers to the list:
L.append(100)
This also demonstrates that the list object has a number of associated methods. Here we used the “append” method. It is like a function that implicitly operates on that element. You can inspect any Python object using the dir() built-in:
dir (L)
This concept is called introspection. From the dir() output, we find a method called insert(). To see what that does, we use an IPython help feature: typing a question mark after an object name prints a small help text:
L.insert?
Here we insert an element in L after element 3:
L.insert(3, "morten")
This illustrates that a list can contain any type of object, in any mixture. Lists can also contain lists:
M=['hej dude'] M.append(L) M.append('hej')
We can access the individual elements of a list using the square bracket notation:
L[1] L[0] L[4]
We can step through (“iterate”) a list in a for loop:
for x in L: print x
1.3.2 Dictionaries
Another very useful built-in compound object is the dictionary. A dictionary is defined using curly brackets:
D = {1:'ditlev', 2:'rune'}
Here, the numbers 1 and 2 are the keys of the dictionary. The strings ‘ditlev’ and ‘rune’ are the values. A key can be any immutable object, like strings:
D['ditlev'] = 'brodersen'
Values of a dictionary can be any object, for example also our list L from before:
D = {1:'ditlev', 2:'rune', 'ditlev':'brodersen', 'mylist':L}
We can iterate through a dictionary in a for loop. Here, we use two dicts ‘isoelectric’ and name that I defined when preparing for this session. (I saved this object earlier using the magic command %store, then it is loaded automatically when IPython starts up.) [Readers of the on-line tutorial can find this and other data in the Appendix.]
for a in isoelectric: print a, name[a], isoelectric[a]
1.3.3 Tuples
Tuples are exactly like lists, except they are immutable (elements can not be modified once defined.) A tuple is defined using parentheses:
T = (42,43,44,2)
Tuples are used extensively internally in Python, for example when passing arguments to functions.
1.3.4 Sets
The set is a very useful Python object. You define it like this:
S = set()
This is an empty set. The set constructor can also take a sequence (e.g. lists, tuples)
S = set([1,2,3,4,5,6])
Sets are unique, an element can exist only once. The set constructor silently discards duplicate elements:
S = set([100,2,2,2,2,23,4,5,6])
The Set() class has an add() method:
S.add("morten")
This illustrates that a set can contain different types of objects. The add() method is similar to the append() method of lists, but “append” doesn’t make sense for a set, since it is unordered.
You can iterate through a set in a for loop:
for i in S: print i
Sets are very powerful. You can use the well-known set operations like union, difference and intersection. Previously, I have defined the set aa containing one-letter codes of the 20 amino acids:
aa = set("ACDEFGHIKLMNPQRSTVWY")
(NOTE: The set aa defined above does not contain the string “ACD…”! To achieve that I would need to use the constructor set([“ACD…”]). That is because the set constructor takes 1 argument which is an iterable object, and the string “ACDE…” iterates to “A”, “C”, …)
We now define sets of the acidic and basic amino acids:
basic = set (('K', 'R')) acidic = set(('D', 'E'))
We can find the set of “neutral” amino acids:
neutral = aa - acidic - basic
The set phobic, which I have defined previously, has the set of hydrophobic amino acids. The hydrophilic set is thus:
philic = aa - phobic
The set of amino acids, that is hydrophilic but not charged is:
philic - (acidic|basic)
The set of amino acids without a side chain charge is:
aa - acidic - basic
1.4 Copying objects
Python is very economical with computer memory, and tries to reference the same memory locations if at all possible. This gives a few unexpected effects that you need to know about. It is normally never any problem.
Simple objects (ints, strings, etc.) can be copied like you expect:
a = 10 b = a
We have assigned a to b, and they are separate, independent objects. If we subsequently alter b, a will stay the same. This is not the case for compound objects, and more complicated objects. For example, create a list:
A = range(10)
(this gives a list of integers 0-9), and make an assignment to B:
B = A
Next, let’s change element 1 of B:
B[1] = 100
You will notice that element 1 of A changes too! So, Python is using the same memory for lists A and B, and B is just another reference to that memory.
We can get around that by using the copy module:
import copy
If we now copy A to B, it will work as expected:
B = copy.copy(A)
B[1] = 200
and A stays the same. However, copy.copy() is a shallow copy. If A contains anything but simple objects (for example another list), we need to use a deep copy. For example, this fails:
C = [A,B] D = copy.copy(C) D[1][1] = 1000 print C
but using the deep copy will work:
D = copy.deepcopy(C) D[1][1] = 2000 print C print D
1.5 Modules
Since we already met the copy module above, let us briefly introduce the concept of modules in Python. In short, a module is a collection of variables, functions and classes that you can load. An often used module is os. It contains a bunch of objects needed if you want to interact with the operating system. Modules need to be imported:
import os
Now, the power of the os module is available to us. You can inspect its content using dir():
dir(os)
Let’s take a look at the uname() object from the os module. This basically does the same information that is given when you type uname from the shell:
x = os.uname() type(x)
You see, that x is a tuple. We can access its elements:
hostname = x[1] print hostname
This was an example of a core module (distributed with Python). There is a very large number of third-party Python packages. One example, that we will study more later, is numpy, Numerical Python.
import numpy dir(numpy)
You can see that Numpy contains a very large number of objects, which all deal with numerical data (“numbers”) in a very efficient way. The most basic object is the array:
a = numpy.array((1,2,3,4,5,6,7,8,9))
This defines a linear array of 9 elements. We can reshape that to a (3,3) array:
a.reshape(3,3)
That array can be recast to a matrix object, and participate in very efficient matrix and vector operations. We will look more into this later.
1.5.1 Namespace
In the above, you have seen that whenever you need to reference an object from a module, you need to prefix it with the name of the module. That is because each module has its own namespace. If you need to reference a specific object a lot, you can import it into the current namespace, like so:
from os import uname
1.6 Functions
Python provides functions. Let us define a function to add two objects:
def mysum(a,b): return a+b
The parenthesis contains the two formal parameters of the function. These exist in the local namespace of the function. Variables a and b in the calling program (if they exist) will not be affected. Let’s test the function:
mysum(10,20) mysum(0.4,20)
This works as expected. However, you can also do this:
mysum('asdfa','aasdasdf')
which demonstrates, that any object providing the method __add__() can be “added” using the ‘+’ operator! However, this:
mysum(0.4,'a')
will raise an exception because floats and strings can’t be added.
1.7 Files
It is very easy to read files from Python. This opens a file and associates a file object f to it:
f = open("1zen.pdb")
Now f is an ordinary Python object that we can do stuff with:
dir(f)
The readlines() method simply swallows the whole file, and returns a list of lines:
L = f.readlines()
Now the lines in the file can be accessed. L[0] contains the first line, and (since this is a PDB file) line 500 is one of the ATOM records:
print L[500]
You can see that L[500] is a string:
type(L[500])
If we want to get to the individual items in that string (the X coordinate, the B factor etc.) we need to split that string into individual components. To do this, we can use the string’s split() method:
m = L[500] mm = m.split()
Now, mm is a list of strings containing the individual items. If we wanted to treat the X-coordinate as a number (say, add 0.01 Angstrom to it) we need to convert it to a number:
x = float(mm[5]) x = x + 0.01
When done with the file, you should close it like a good boy:
f.close()
Files can be iterated like we saw for lists:
L = [] f = open("1zen.pdb") for line in f: L.append(line.split())
Here, we did several of the steps above in one operation.
1.8 File-like objects
We saw that Python’s file object is very convenient, so Python operates with so-called file-like objects. These are objects that can be anything, but appear to the programmer as though they are simple files. A simple example shows that you can access a web page as though it was a file:
import urllib f = urllib.urlopen('http://www.bioxray.au.dk/index_uk.php')
Now we have our “file-like” object f, and we can suck in the web page.
W = f.readlines() print len
We get a list of 413 lines containing the HTML text. However, the “file-like” object f can do more than an ordinary file. For example, you can get the HTTP header information:
print f.info()
but that is another story 🙂
1.9 Take-home message
The take-home message of this presentation is:
In Python, EVERYTHING IS AN OBJECT
Appendix
# Amino acid properties hydropathy = {'A': 1.8, 'C': 2.5, 'D': -3.5, 'E': -3.5, 'F': 2.8, 'G': -0.4, 'H': -3.2, 'I': 4.5, 'K': -3.9, 'L': 3.8, 'M': 1.9, 'N': -3.5, 'P': -1.6, 'Q': -3.5, 'R': -4.5, 'S': -0.8, 'T': -0.7, 'V': 4.2, 'W': -0.9, 'Y': -1.3} tlc = {'A': "ALA", 'C': "CYS", 'D': "ASP", 'E': "GLU", 'F': "PHE", 'G': "GLY", 'H': "HIS", 'I': "ILE", 'K': "LYS", 'L': "LEU", 'M': "MET", 'N': "ASN", 'P': "PRO", 'Q': "GLN", 'R': "ARG", 'S': "SER", 'T': "THR", 'V': "VAL", 'W': "TRP", 'Y': "TYR"} charge={'A': 0, 'C': 0, 'D': -1, 'E': -1, 'F': 0, 'G': 0, 'H': 0.1, 'I': 0, 'K': 1, 'L': 0, 'M': 0, 'N': 0, 'P': 0, 'Q': 0, 'R': 1, 'S': 0, 'T': 0, 'V': 0, 'W': 0, 'Y': 0} isoelectric={'A': 6.0, 'C': 5.02, 'D': 2.77, 'E': 3.22, 'F': 5.48, 'G': 5.97, 'H': 7.47, 'I': 5.94, 'K': 9.59, 'L': 5.98, 'M': 5.74, 'N': 5.41, 'P': 6.3, 'Q': 5.65, 'R': 11.15, 'S': 5.68, 'T': 5.64, 'V': 5.96, 'W': 5.89, 'Y': 5.66} mw = {'A': 89.09, 'C': 121.15, 'D': 133.1, 'E': 147.13, 'F': 165.19, 'G': 75.07, 'H': 155.16, 'I': 131.17, 'K': 146.19, 'L': 131.17, 'M': 149.21, 'N': 132.12, 'P': 115.13, 'Q': 146.15, 'R': 174.2, 'S': 105.09, 'T': 119.12, 'V': 117.15, 'W': 204.23, 'Y': 181.19} name = {'A': 'alanine', 'C': 'cysteine', 'D': 'aspartic acid', 'E':'glutamic acid', 'F': 'phenylalanine', 'G': 'glycine', 'H': 'histidine', 'I': 'isoleucine', 'K': 'lysine', 'L': 'leucine', 'M': 'methionine', 'N': 'asparagine', 'P': 'proline', 'Q': 'glutamine', 'R': 'arginine', 'S': 'serine', 'T': 'threonine', 'V': 'valine', 'W': 'tryptophan', 'Y': 'tyrosine'} polarity={'A': 'n', 'C': 'n', 'D': 'p', 'E': 'p', 'F': 'n', 'G': 'n', 'H': 'p', 'I': 'n', 'K': 'p', 'L': 'n', 'M': 'n', 'N': 'p', 'P': 'n', 'Q': 'p', 'R': 'n', 'S': 'p', 'T': 'p', 'V': 'n', 'W': 'n', 'Y': 'p'} aa = set("ACDEFGHIKLMNPQRSTVWY") phobic = set([ x for x in hydropathy if hydropathy[x] > 0.1]) neutral = set("ANCEGILMFQPSTWYV") positive = set("RK") negative = set("DE") zwitter=set("H")
Date: 2012-01-26 11:22:16 CET
HTML generated by org-mode 6.21b in emacs 23
Disk Quota Exceeded?
The disk quota system in Linux is quite old, and if you have a distributed workstation environment with NFS mounted shares it is not possible in a convenient way to inform users that they have exceeded their disk limits.
So we created a simple disk quota warning system, written in Python, that works using the notification interface (libnotify). A user who has exceeded the disk quota is then at regular intervals presented with a pop-up warning like the one shown in the figure below.
The quotawarn system is divided into three parts. Two of them run on the file server, and one on the users workstation, under the users’ own UID.
On the file server, there is a program called quotawarn.cron, which (duh!) should be run at regular intervals by cron. This program simply runs repquota and squirrels away the information in a convenient format. Currently, this is simply a pickled Python dictionary indexed by UID, but could of course also be a fancier database storing historical data on each user’s disk usage.
The second program on the server is a CGI script. Using the standard web interface lets us rely on the access settings in the HTTP server, thus minimizing the need to program a special security system, and relieving the burden on the sysadm of learning and maintaining yet another access control system.
Quotawarn.cgi responds to messages from the network, replying to requests for specific UIDs by transferring the data for that user in JSON format. Or, optionally, by creating an HTML page with the data of all users.
On each workstation, the notification script quotawarn is started when the user logs on. The sysadm can achieve this by placing an entry in /usr/share/autostart. The program will ask for data from quotawarn.cgi on the file server, and if the user is under the disk quota limit, the program will exit without any more noise. If, however, the users disk quota is exceeded, the program will pop up a notification (shown below). This will continue at regular intervals (set by the sysadm).
A second notification system, yet to be developed, could hook into Ubuntu’s motd system, and provide the necessary information when the user logs on via an ssh connection to the network.
I have created a project on Launchpad for quotawarn in the hope that others would like to contribute to the project, and make it more generally usable. Although an attempt has been made to make the system general, we have not been able to test it in other settings than our own. So, if this has whet your appetite, please feel free to branch the project and contribute!
Free and completely Ubuntu (2)
The other day, I reported here that Tuesday evening (Feb. 8.), the popular tech-review program on danish public television channel DR2 “So Ein Ding…” dedicated an entire episode to Ubuntu.
A commenter asked for a translation, so here it is, including the program host, Nicolaj Sonne’s evocative sound effects.
Hey!
Today we jump in, and go all-in with Open Source. I have purchased a compltely new Low-budget computer, nuked Windows, and installed the free OS Ubuntu. In other words, I don’t need to spend one more Krone on this computer!
Welcome to this episode of “So ein Ding”!
…
1:35 As always, we are playing with open cards, and this part will be a bit different “Ding” than usual. The thing is, a computer OS is a seriously complicated thing. Imagine you arrive in a new city. It will not take long before you can find your way around, but it will take a few years before you know every sneaky little watering-hole from Christianshavn to Frederiksberg. And to stay within that picture, there are three different cities in the land of Operating Systems: Windows, Mac OS and Linux.
My first PC was a 486DX. About 15 years ago, I installed Windows on it, and I have used that ever since. Not on an “ultra-user” level, buuuut… I have re-installed one driver or another now and then.
1:42 Then about five years ago, I was forced to re-school in another OS, namely Mac OS. I got a new boss, who told me, “hey, here in the firm, we use computers with apples on, period.”
About two weeks ago, I started using Ubuntu. I have used Ubuntu before, and other Linux versions in the same family as Ubuntu, but it was rather sporadic.
So you can rightfully say, I am a completely green Ubuntologue, and it it these experiences we are going to talk about today.
2:17 So now, Ladies and Gentlemen, we present the danish sneak-preview on “This is how you install an operating system”. Perhaps it will get your cold-sweat running down your back and your eyes are flickering, when words like: “boot”, “partition”, “NTFS”, “ext4”, and so on, fly through the air. But hang on, because if you do that, you will be free-wheeling software-wise in the all foreseeable future.
You go to Ubuntu.com, press “download” and then you fetch a file called “an image”, burn it to a CD or put it on a USB-stick.
And already now, it is obvious to see that they want customers in the shop, because there is a super, super easy step-by-step guide. Just follow that!
Well, I did not need the pre-installed Windows version, so ummphh, it got the knife.
Regarding the installation, it was around 15 minutes, click, click, click, a cup of coffee, and then, everything worked. All key-combos, turn the backlighting up and down, Wi-Fi on/off, sound, and things like that. Everything apparently worked, it was fantastic!
THEN the system proposed an update, and, from old habit, YES! I want that update. It took around 10 minutes, and then I came back to this: a computer, where neither mouse nor keyboard works!
No reaction what-so-ever. Then I thought, perhaps, if I attach an external USB mouse and USB keyboard. That worked, then I could google a bit around, and I discovered, I was not the only one losing contact with the mouse and keyboard.
Well, that was a bit of a burn-out. The thing is, I had fetched — as usual, from habit — the newest 10.10 version of Ubuntu. Then I tried with Ubuntu 10.04 LTS — Long Term Support — you can read about it on Ubuntu’s home-page, it makes sense. I installed that, and that worked perfectly, updates and everything. Except ONE small thing, and that was: I had sound though the phone-jack, but not through the speakers!
IF your speakers don’t work but your phone-jack does, then you “just” need to open a terminal window, and write “sudo apt-get repository blah blah blah”.
You need to be good at googling, and you need to have a bit of courage. It is not rocket science but it is not something I would want to explain to my mother over the phone, aaaand: my mother is a programmer. Well, a programmer from the days when the IBM 1287 was the latest & greatest, and programs was something you wrote by hand, and then sent out to be punched by “punch-ladies” who would… punch your programs
The free OS Ubuntu is a part of the Linux family which is open source software. Ubuntu means, loosely translated: “I am who I am because of who we all are.” What a bunch of hippie smoke! 😉
But it is not all psycodelic, there is a meaning to the madness, Ubuntu is one of the more well-known open-source software projects, and in brief, “open source” means that the “recipe”, the source, is freely available. This is NOT true for Mac Os and Windows. There, the engine room is shut and locked!
Even though Ubuntu is entirely free, it is presumably neither security-wise or economically completely out in the woods, because e.g. the French parliament, the French police, and the Swedish police all use Ubuntu.
Ubuntu is, as mentioned, a part of the Linux family. “Linux what-creature?”. Well, you meet Linux really often! Six out of ten times you write http://www.some-homepage.dk, there is a Linux server in the other end.
6:40 And now, the Ubuntu-sun rises! [shows the Ubuntu install ID like a rising sun] I now have a computer, completely without pre-installed Windows, where everything works!
Here, right in the middle of the graphical user interface, you will relatively quickly feel at home. Things are organized a bit in-between Windows and MacOSX But, with respect to the tightness of things, where applications are found, files, folder-structure and so on, it is probably more like Mac OSX.
But, what about compatibility? Well, those days, where computers could not speak to each other… it’s no longer like that. You can easily create a file on a Ubuntu computer, put that file on a USB-stick, put that into a Mac, burn a CD, and then read that CD on a Windows machine. It is not a problem. So yes, you can easily send a mail to your grandma, with holidays pictures, music and video. Even if your grandma uses Ubuntu, everything will work just fine.
There IS lots of software, that only works for e.g. Windows or Mac OS, and if you save a fike in that kind of software, then you cannot open it in Ubuntu… at least it is not certain that you can.
8:00 Tjjj , tjjj, tjj, tjj [plays around with the UI] … In spite of fancy animations, there are some things I don’t think are so polished in Ubuntu as they are under Windows and Mac OS, and an example is navigating between programs with keystrokes while you are grabbing something with the mouse. If we look at an example here: Here in my Mail program. I say; I want to make a mail, wwwyycchh, and then we write a mail, write-write-write. Then I realize, I would like to attach a photo to that mail. Then I use my key-strokes to get to my pictures. Then we look at it… here is a funny picture, we want to attach that. Then, aaannnchh, you can grab it with the mouse. On a Windows computer, on a Mac, I would be able to — while I’ve grabbed this image — to use key-combos to get back to the mail program… I can’t do that here! And it’s really irritating!
Over here, on my typewriter, it doesn’t run Windows, it runs Mac OS, I can search files in a neat way. If I want to find all the photos I have taken with a Nokia N73 phone (jpeg because it is pictures)… in no time, all the photos I have taken with that phone, they are here! That is very smart, because this information is not something associated with the file-name, it is something inside the file itself that I can search. I can’t do that in Ubuntu, there isn’t that kind of meta-meta-meta search. That is something I miss enormously!
One of the things that make Ubuntu somewhat special is that it has a built-in appstore, a bit like you know it from the mobile-phone world: small programs you can fetch. It is called “Ubuntu Software Center” and it is loaded with software! Right now there are 32648 different items, that all are 100% verified and totally OK programs.
Then you can of course browse around after categories, but now, I would like to fetch a program that can make DVDs, that is DVD graphics and so on…. that’s called
DVD authoring. Let’s see [browses the appstore] … wwyyyccchh, “DVD-ripper”.. “trans”… bmm, bmm… “auto”… eetchhh… “Author DVDs and slideshow” … Then I can click here, select “More Info”, then you can read a bit about what the program can do… it looks good… “Install”… and its free. Then it asks me for my password…. mmmwrrcchh…
And that’s every time I want to alter something on the machine, then it asks for my password, which I created during the installation. Now it starts installing…. dddlidstchh…
In the meantime, we can take a look, over here, on “Installed Software”, that is also shown in one place, and here I can see all the software that’s installed on the computer. And if I feel like… wwyyycchhh… pull the plug on one of them, I just click on “Remove”. No nonense.
And now, my DVD program should be installed: dlllyytchh boing! And, now, get started makeing DVDs!
But, take it easy, you are not forced to only “purchase” in Ubuntu’s Software Center. You can use everything you can find on the net. Of course, it must generally be made for Ubuntu… and note I say “generally”, because the thing is, I’ve fetched this little thing here, it’s called “Wine”. And then you can see here… Wine means “Wine Is Not an Emulator”… and then you can install Windows programs… wwyyyytccchhh. Now I’ve fetched “Picasa” in the Windows version, and it now runs under Ubuntu. It is not certain that exactly your favourite program runs under this “Wine”, but then perhaps you can find another favourite program that does.
With Ubuntu I can use my netbank. But its €&&#%#% no thanks to NemId [danish net id organization], who absolutely offers NO support to Linux. Definitely! (Well. I’ll keep an eye on you!)
Then I can of course burn CDs DVD, i can import photos from my digital camera, I can write mails, I can create documents — it’s not called “Word”, it’s called “OpenOffice” — but I can by and large do everything I usually do. And naturally, all the things I do on the net, I can do here. Even this fellow, a 3G modem, no problems. I put it in the USB port, Yes”, “Yes” and “Yes Sir” and then I was on the net in… 1.5 minute.
Perhaps I have been damned lucky with Ubuntu… perhaps not.
12:30 Very fine and very very free, but with that said, it must be said, before the Ubuntu-sun sets, that compared to the dominating, pre-installed alternative, I can’t really see the great advantage, except perhaps economy and ideology. And! You don’t need to bother with these… [several dialogue boxes appear “Your TV can break down anytime”]. The final verdict is at the end of the program.
[Ubuntu gets 4/6]
Free and completely Ubuntu
Tuesday evening (Feb. 8.), the popular tech-review program on danish public television channel DR2 “So Ein Ding…” dedicated an entire episode to Ubuntu. The program host, Nikolaj Sonne, took Ubunto for a test spin, and gives a very positive, fair and balanced review. His final verdict ends on 4 out of 6, which is extremely good, especially considering that an upgrade broke his Maverick installation in the trials (no trackpad, no keyboard) and he had to downgrade to Lucid.
You can view the episode here — in danish of course — but watch the program anyway for the nice graphics.
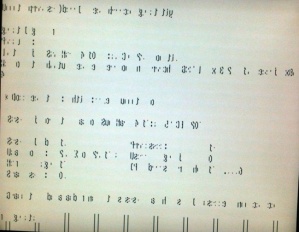


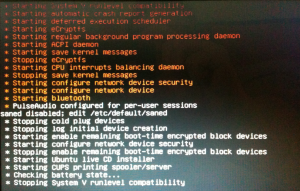


26 comments